Kernel density estimation for circular functions
28th March, 2014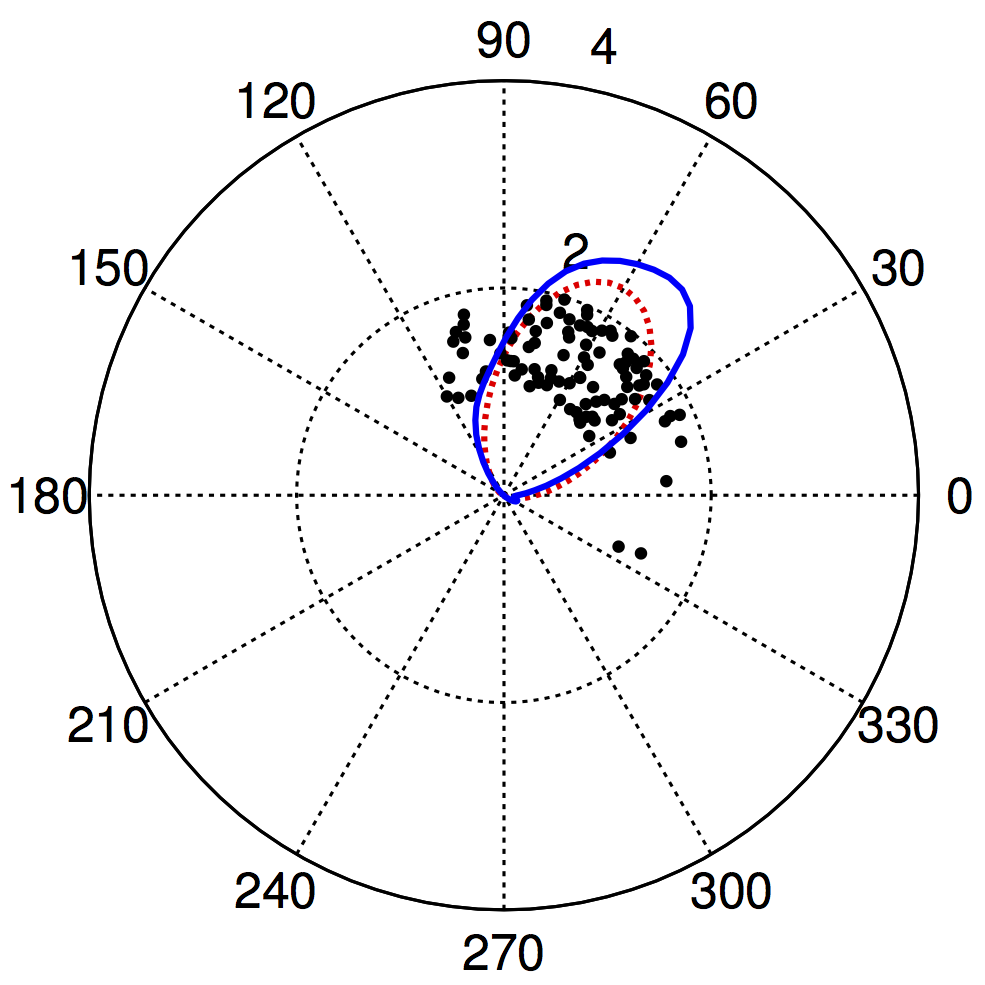
Matlab code for performing kernel density estimates on periodic and aperiodic domains, for unidimensional and multidimensional data.
Estimating multi-dimensional PDFs over periodic domains is reasonably straightforward, but can't be done with Matlab's built-in tools. Here are two functions for kernel density estimates for circular functions. By default they will estimate a PDF over the periodic domain, and can be used with weighted and unweighted data.
Download and installation
Download the source code: circ_ksdensity.zip
Unzip the file into a directory on the matlab path. Two functions are included: circ_ksdensity and circ_ksdensityn.
Uni-dimensional kernel density estimates — circ_ksdensity
Usage:
[vfEstimate] = circ_ksdensity(vfObservations, vfPDFSamples, <vfDomain, fSigma, vfWeights>)
This function calculates a kernel density estimate of an (optionally weighted) data sample, over a periodic domain.
vfObservations is a set of observations made over a periodic domain, optionally defined by vfDomain: [fMin fMax]. The default domain is [0..2*pi]. vfPDFSamples defines the sample points over which to perform the kernel density estimate, over the same domain as vfObservations.
Weighted estimations can be performed by providing the optional argument vfWeights, where each element in vfWeights corresponds to the matching element in vfObservations.
The kernel density estimate will be performed using a wrapped Gaussian kernel, with a width estimated as
(4/3)^0.2 * circ_std(vfObservations, vfWeights) *(length(vfObservations^-0.2)
The optional argument fSigma can be provided to set the width of the kernel.
vfEstimate will be a vector with a (weighted) estimate of the underlying distribution, with an entry for each element of vfPDFSamples. If no weighting is supplied, the estimate will be scaled such that it forms a PDF estimate over the supplied sample domain, taking into account sample bin widths. If a weight vector is supplied then the estimate will be scaled such that the sum over the domain attempts to match the sum of weights, taking into account sample bin widths.
Multi-dimensional kernel density estimates — circ_ksdensityn
Usage:[vfEstimate, vfBinVol] = circ_ksdensityn(mfObservations, mfPDFSamples, <mfDomains, vfSigmas, vfWeights>)
This function calculates a kernel density estimate of an (optionally weighted) data sample, over periodic and aperiodic domains. The sample is assumed to be independent across dimensions; i.e. density estimation is performed independently for each dimension of the data.
mfObservations is a set of observations made over a (possibly periodic) domain. Each row corresponds to a single observation, each column corresponds to a particular dimension. By default all dimensions are periodic in [0..2*pi]; this can be modified by providing the optional argument mfDomains. Each row in mfDomains is [fMin fMax], one row for each dimension in mfObservations. If a particular dimension should not be periodic, the corresponding row should be [nan nan]. Bounded support over a dimension is NOT implemented; each dimension is either linear and infinite or periodic.
mfPDFSamples defines the sample points over which to perform the kernel density estimate, over the same domains as mfObservations.
Weighted estimations can be performed by providing the optional argument vfWeights, where each element in vfWeights corresponds to the matching observation in mfObservations.
The kernel density estimate will be performed using a multivariate Gaussian kernel, independent along each dimension, and wrapped along the periodic dimensions as appropriate. Kenel widths over periodic dimensions are estimated as
(4/3)^0.2 * circ_std(mfObservations(:, nDim), vfWeights) * (length(mfObservations)^-0.2)
Kernel widths over non-periodic dimensions are estimated as
(4 * std(mfObservations(:, nDim), vfWeights)^5 / 3 / length(mfObservations))^(1/5)
The optional argument vfSigmas can be provided to set the width of each kernel.
vfEstimate will be a vector with a (weighted) histogram estimate of the underlying distribution, with an entry for each point in mfPDFSamples. If no weighting is supplied, the estimate will be scaled to estimate a PDF over the supplied multi-dimensional domain, taking into account the estimated volume of each bin. If a weight vector is supplied, the estimate will be scaled such that the sum over the domain attempts to match the sum of weights, taking into account the multi-dimensional bin volumes.
vfBinVol is a vector containing volume estimates for each row in mfPDFSamples, under the assumption that each dimension is linearly scaled and mutually orthogonal.